UHerding#
- class skactiveml.pool.UHerding(method='margin_sampling', predict_proba_dict=None, predict_proba_parser=None, temperatures=None, validation_size=0.2, n_ece_bins=15, normalize_samples=True, metric='rbf', metric_dict=None, adaptive_sigma=True, missing_label=nan, random_state=None)[source]#
Bases:
SingleAnnotatorPoolQueryStrategyUncertainty Herding (UHerding)
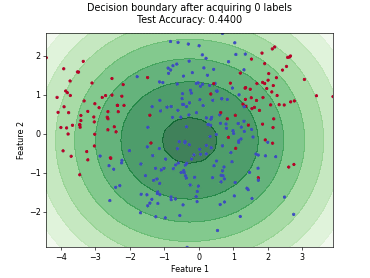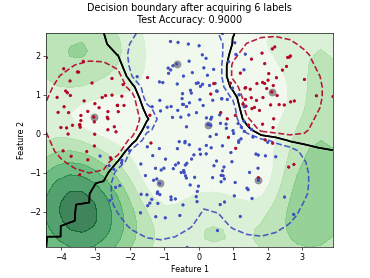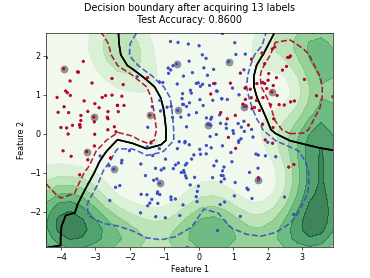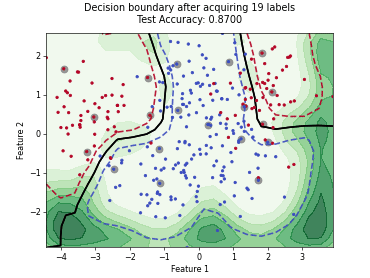
“Uncertainty Herding” (UHerding) is a query strategy [1] that greedily maximizes an uncertainty-weighted coverage objective in feature space. In addition to the greedy selection itself, the implementation follows the parameter adaptation scheme of the paper:
select a temperature based on calibration via train/validation splits of the currently labeled set,
adapt the Gaussian kernel radius to the current labeled feature space.
- Parameters:
- method‘least_confident’ or ‘margin_sampling’ or ‘entropy’, default=’margin_sampling’
Uncertainty definition applied to temperature-scaled probabilities.
- predict_proba_dictdict or None, default=None
Optional keyword arguments forwarded to clf.predict_proba to request additional outputs such as logits and embeddings.
If predict_proba_parser is None, optional outputs are interpreted by the default convention (probas, logits, embeddings). Typical usage with SkorchClassifier is therefore:
predict_proba_dict={"extra_outputs": ["logits", "emb"]}
If logits are not returned by predict_proba, decision_function is used as a fallback when available, e.g. for scikit-learn logistic regression models wrapped by SklearnClassifier.
- predict_proba_parsercallable or None, default=None
Optional parser applied to the raw return value of clf.predict_proba(X, **predict_proba_dict).
The parser must return either (probas, logits) or (probas, logits, embeddings). probas may be None, in which case they are computed from logits via softmax. embeddings may be None, in which case the original samples are used.
If None, the default convention is used:
array output: treated as probas,
tuple output: treated as (probas, logits, embeddings).
- temperaturesfloat or array-like of shape (n_temperatures,) or None, default=None
Candidate temperatures used during the calibration search. If a single positive float or a length-one array is provided, that temperature is used directly without internal calibration refits. If None, temperatures=np.logspace(-1, 1, 49) is used.
- validation_sizefloat or int, default=0.2
Validation size passed to the calibration train/validation split.
- n_ece_binsint, default=15
Number of bins used for the expected calibration error.
- normalize_samplesbool, default=True
Flag whether to normalize feature vectors to unit length before computing pairwise distances and kernels.
- metricstr or callable, default=’rbf’
Kernel used for the coverage objective.
- metric_dictdict or None, default=None
Optional keyword arguments passed to pairwise_kernels.
- adaptive_sigmabool, default=True
Flag whether to adapt the radius according to the minimum non-zero labeled pairwise distance. This option requires metric=’rbf’.
- missing_labelscalar or string or np.nan or None, default=np.nan
Value to represent a missing label.
- random_stateNone or int or np.random.RandomState, default=None
The random state to use.
References
[1]W. Bae, G. Oliveira, and D. J. Sutherland. “Uncertainty Herding: One Active Learning Method for All Label Budgets.” In Int. Conf. Learn. Represent., 2025.
Methods
Get metadata routing of this object.
get_params([deep])Get parameters for this estimator.
query(X, y, clf[, fit_clf, sample_weight, ...])Determines for which candidate samples labels are to be queried.
set_params(**params)Set the parameters of this estimator.
- get_metadata_routing()#
Get metadata routing of this object.
Please check User Guide on how the routing mechanism works.
- Returns:
- routingMetadataRequest
A
MetadataRequestencapsulating routing information.
- get_params(deep=True)#
Get parameters for this estimator.
- Parameters:
- deepbool, default=True
If True, will return the parameters for this estimator and contained subobjects that are estimators.
- Returns:
- paramsdict
Parameter names mapped to their values.
- query(X, y, clf, fit_clf=True, sample_weight=None, candidates=None, batch_size=1, return_utilities=False)[source]#
Determines for which candidate samples labels are to be queried.
- Parameters:
- Xarray-like of shape (n_samples, …)
Training data set, usually complete, i.e., including the labeled and unlabeled samples.
- yarray-like of shape (n_samples,)
Labels of the training data set (possibly including unlabeled ones indicated by self.missing_label).
- clfskactiveml.base.SkactivemlClassifier
Classifier implementing fit and predict_proba. For temperature-scaled uncertainty estimation, the classifier should either provide logits via predict_proba extras or implement decision_function. Otherwise, the non-calibrated probabilities are used as fallback.
- fit_clfbool, default=True
Defines whether the classifier clf should be fitted on X, y, and sample_weight before evaluating the acquisition function. Independent of this flag, temporary cloned classifiers may still be fitted internally to select the temperature parameter.
- sample_weightarray-like of shape (n_samples,), default=None
Weights of training samples in X.
- candidatesNone or array-like of shape (n_candidates,), dtype=int or array-like of shape (n_candidates, …), default=None
If candidates is None, the unlabeled samples from (X, y) are considered as candidates.
If candidates is of shape (n_candidates,) and of type int, candidates is considered as the indices of the samples in (X, y).
If candidates is of shape (n_candidates, …), the candidate samples are directly given in candidates (not necessarily contained in X).
- batch_sizeint, default=1
The number of samples to be selected in one AL cycle.
- return_utilitiesbool, default=False
If True, also return the utilities based on the query strategy.
- Returns:
- query_indicesnumpy.ndarray of shape (batch_size,)
The query indices indicate for which candidate sample a label is to be queried, e.g., query_indices[0] indicates the first selected sample.
If candidates is None or of shape (n_candidates,), the indexing refers to the samples in X.
If candidates is of shape (n_candidates, …), the indexing refers to the samples in candidates.
- utilitiesnumpy.ndarray of shape (batch_size, n_samples) or numpy.ndarray of shape (batch_size, n_candidates)
The utilities of samples after each selected sample of the batch, e.g., utilities[0] indicates the utilities used for selecting the first sample (with index query_indices[0]) of the batch. Utilities for labeled samples or already selected candidates are set to np.nan.
If candidates is None, the indexing refers to the samples in X.
If candidates is of shape (n_candidates,) and of type int, utilities refers to the samples in X.
If candidates is of shape (n_candidates, …), utilities refers to the indexing in candidates.
- set_params(**params)#
Set the parameters of this estimator.
The method works on simple estimators as well as on nested objects (such as
Pipeline). The latter have parameters of the form<component>__<parameter>so that it’s possible to update each component of a nested object.- Parameters:
- **paramsdict
Estimator parameters.
- Returns:
- selfestimator instance
Estimator instance.